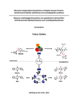
| Titel: | Ribosome-Independent Biosynthesis of Peptide Natural Products: Nonribosomal Peptide Synthetases and Cyclodipeptide Synthases |
| Autor: | Gießen, Tobias |
| Weitere Beteiligte: | Marahiel, Mohamed A. (Prof. Dr.) |
| Veröffentlicht: | 2013 |
| URI: | https://archiv.ub.uni-marburg.de/diss/z2013/0411 |
| URN: | urn:nbn:de:hebis:04-z2013-04115 |
| DOI: | https://doi.org/10.17192/z2013.0411 |
| DDC: | Chemie |
| Titel (trans.): | Ribosom-unabhängige Biosynthese von peptidischen Naturstoffen: Nichtribosomale Peptidsynthetasen und Cyclodipeptidsynthasen |
| Publikationsdatum: | 2014-02-24 |
| Lizenz: | https://rightsstatements.org/vocab/InC-NC/1.0/ |
| Schlagwörter: |
|---|
| cyclodipeptide synthases, Ribosom-unabhängige Peptidsynthese, Cyclodipeptide, Naturstoffchemie, Peptide, Diketopiperazine, Antibiotikum, diketopiperazines, Nonribosomal peptides, Nichtribosomale Peptide, nonribosomal peptide synthetases, Molekularbiologie, Biochemie, Synthetische Biologie |
Summary:
Bioactive peptide natural products continue to play an important role in modern medicine since many of them represent last-resort treatments for a variety of life threatening diseases. A large fraction of those peptides are generated via ribosome-independent templated and non-templated biosynthetic pathways. Oftentimes the structural complexity of those peptides has prevented their synthesis and diversification through purely synthetic means. New approaches build on detailed knowledge about the respective biosynthetic pathways have been envisaged where fermentative in vivo or chemoenzymatic strategies are used to generate highly modified peptide natural products. To work towards this vision two templated and two non-templated biosynthetic pathways have been investigated. Both their abilities to generate particular peptide scaffolds and to subsequently decorate a given peptide backbone through the action of dedicated modification enzymes have been explored.
The two templated NRPS-dependent pathways studied are the biosynthetic systems of the antitumor-antibiotic sibiromycin and the newly discovered siderophore mirubactin.
The origin of the highly substituted anthranilate moiety found in sibiromycin was investigated. The pathway was shown to consist of four steps starting from the known metabolite 3-hydroxykynurenine using detailed in vitro analyses. Initially, the SAM-dependent methyltransferase SibL converts its substrate to the 4-methyl derivative, followed by hydrolysis through the PLP-dependent kynureninase SibQ leading to 3-hydroxy-4-methylanthranilic acid (3H4MAA) formation. Then the NRPS didomain SibE activates 3H4MAA and tethers it to its thiolation domain, where it then serves as the hydroxylation substrate for the FAD/NADH-dependent hydroxylase SibG, yielding the fully substituted anthranilate moiety found in sibiromycin.
The siderophore mirubactin was discovered and purified from Actinosynnema mirum through cultivation under iron-limited conditions followed by activity-guided isolation. Structure elucidation was accomplished through a combination of spectroscopic, mass spectrometric and derivatization methods. Bioinformatic analyses coupled with in vitro characterization of its biosynthetic machinery, was used to identify the mirubactin gene cluster. A biosynthetic assembly route could be proposed comprising the iterative use of a stand-alone carrier-protein-bound substrate (MrbD) and the formation of an unusual O-acyl hydroxamic acid ester bond through a C-terminal condensation domain (MrbJ).
In addition, two non-templated CDPS-dependent pathways responsible for nocazine biosynthesis and the generation of singly and doubly methylated ditryptophan diketopiperazines (DKPs) have been investigated.
The first nocazine gene cluster could be identified through bioinformatic analyses and the biosynthetic pathway leading to the two nocazine family members nocazine E and XR334 could be elucidated through in vivo and in vitro studies. DKP-formation is carried out by a CDPS (Ndas_1148) that shows an unknown product profile forming cyclo(L-Phe-L-Tyr) and cyclo(L-Phe-L-Phe) as its main products. Tailoring of the DKP-scaffold is achieved through the combined and combinatorial action of a cyclic dipeptide oxidase (Ndas_1146/1147) and two distinct SAM-dependent O-/N-methyltransferases (Ndas_1145 and Ndas_1149).
A CDPS gene cluster responsible for the formation of methylated ditryptophan DKPs was identified in A. mirum through bioinformatic genome analysis. The assembly pathway was investigated through in vivo and in vitro studies. Initially, the highly specific CDPS Amir_4627 catalyzes the formation of a formerly unknown CDPS product, namely cyclo(L-Trp-L-Trp) followed by the methylation of one or both DKP-ring nitrogens through the action of the SAM-dependent N-methyltransferase Amir_4628 generating singly and doubly methylated ditryptophan DKPs.
 | Das Dokument ist im Internet frei zugänglich - Hinweise zu den Nutzungsrechten |